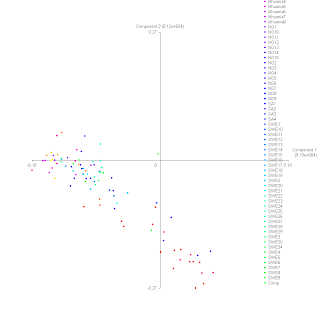
We continue the analysis of the ancient Gotlanders vs our project participants.
In the last analysis using ADMIXTURE we assumed that all SNP's where independent from each other. However in reality we are working with segments of linked SNP's or haplotypes. Chromopainter can analyse haplotype and Finestructure can build a tree based on these.
Input: 22k SNP's. MCMC: 100 000, 100 000, Tree: 10 000, 100 000. HapMap recombination map. c-factor: 0.0262 (effective number of independent genetic element).
THE RESULTS:
FIRST ANALYSIS:
ChunkCounts:
The number of shared segments tree appears to set the ancient Gotlanders at its own branch already at the root of the tree. The closest neighbour is the common Finnish and Saami branch. The branching seem to indicate that Finns and Saamis are closer to the ancient Gotlanders than other populations.
The Finnish cluster is closer maybe indicate a closer relationship than the Saami. It would then be in accordance with the earlier ADMIXTURE analysis:
Note that the the various degree of sharing to the ancient Gotlanders at the rightmost vertical partly yellow strip. Yellow indicate more distant relationship gradiant towards blue is closer relationship (see vertical bar to the right of the matrix).
The ancient Gotlanders appears to have various degree of sharing to other individuals. The reason why the tree is so fragmentet is because of the very low c-factor. It indicate that the elemets are very fragmentet because of few markers (22k). A c-factor of 0 is basically independent SNP's while a c-factor of 1 is basically linked SNP's. In our last Finestructure analysis this factor using 289k was a round 0.48
The total lenght of shared segments indicate much the same structure as mentioned above but with some internal movement.
EXPERIMENTING:
I did however during my own experiments with Finestructure play around with the c-factor parameter. If I set it to 1 meaning considering the result as one independent genetic element the results changed.
ChunkCounts:
We can now also see from the coloring vs the ancient Gotlanders that its only the two Saami groups that have increased sharing with the ancient Gotlanders at the rightmost vertical that move into pink meaning the closest.
ChunkLenght:
The total segments shared give a similar story but all the Saamis are now into the same cluster. The coloring seem to indicate that Saamis have the closest relationship, then Scandinavians, then Lithuanians and then finally Finns. This is not in accordance with the earlier ADMIXTURE analysis.
The first analysis appears to be in accordance with ADMIXTURE and is after the book prescribed by the authors of Finestructure. Finns are the closest to the ancient Gotlanders when assuming many genetic elements,
However if assuming only one genetic element the result is in accordance with Chromopainter raw output data for ChunkCounts and ChunkLenght that says that on population average Saamis are the closest to the ancient Gotlanders, then Scandinavians, then Lithuanians and finally Finns.
In my earlier analysis below using only Chromopainter without the help of Finestructure the population assigment for individuals have been correct for most individuals and this analysis appears to show correct and known relationships between the populations in question when assuming c-factor of 1. If assuming a c-factor of 0 the analysis would probably converge more with the ADMIXTURE analysis.
It seems therefor to me that the clustering of Finns close to the Saami maybe be due to more recent connections like migration of Finns to Saami. This because the connection to the ancient Gotlander appears more distant for Finns than for Saami.
The connection of the ancient Gotlander to the Scandinavians seems to be direct. It doesnt seem to have gone through the North-Saamis or Finns as they look like today to Scandinavians. However we know little about the genetics of the South-Saamis. The individuals of partly South-Saami ancestry that show up in the MDS-plots have not maanged to differentiate themself enough in this analysis from Scandinavians.
(Updated 01/07/2012)